Last updated: 19 February 2009.
Features
- Multi-view 3D graphics
- Stereoscopic viewing using CrystalEyes goggles (or equivalent) is supported on both SGI Irix and Intel Linux.
- Integration of sequence viewing and alignment editing with 3D molecular graphics
- A hierachichal tree view to chemical data
- Molecular surface calculation and visualization
- Visualization of electron density and Grid / AutoDock / DelPhi maps
- Fully automatic protein structure superposition using a C-alpha packing profile correlation
- Modular architecture: dynamically loaded plugin modules, application programming interface based on standard C++
Screenshots
Click on the images to view the full-size versions.
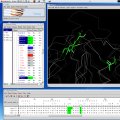 Bodil has three kinds of views to molecular data: a hierarchical tree
view (left), three-dimensional molecular graphics (right) and a
sequence alignment view (bottom). All views show the same
objects. When e.g. a column of the sequence alignment is selected, the
corresponding residues of each structure in the alignment become
selected (green) in the molecular graphics view, too.
Bodil has three kinds of views to molecular data: a hierarchical tree
view (left), three-dimensional molecular graphics (right) and a
sequence alignment view (bottom). All views show the same
objects. When e.g. a column of the sequence alignment is selected, the
corresponding residues of each structure in the alignment become
selected (green) in the molecular graphics view, too.
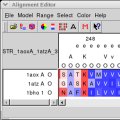 Sequence alignments can be edited interactively and coloured using
predefined or user-supplied colouring schemes. The screenshot shows
colouring according to hydrophobicity.
Sequence alignments can be edited interactively and coloured using
predefined or user-supplied colouring schemes. The screenshot shows
colouring according to hydrophobicity.
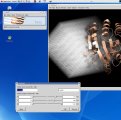 Grid objects (density maps, Grid maps, molecular surfaces etc.) can be
cropped using an interactive tool.
Grid objects (density maps, Grid maps, molecular surfaces etc.) can be
cropped using an interactive tool.
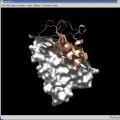 Bodil can display opaque and semitransparent molecular surfaces.
Bodil can display opaque and semitransparent molecular surfaces.